How is everyone not talking about this paper?
It's about a superantigen insert in the spike of SARS-2 that isn't in the spike of SARS.
Update: These guys say there’s no superantigen site in the spike protein as described in this recent publication entitled: “SARS-CoV-2 Spike Does Not Possess Intrinsic Superantigen-like Inflammatory Activity”.1
This Substack is about another danger associated with the spike protein, and not just the viral spike. This danger is called a superantigen. A superantigen can activate all sorts of T cells non-specifically to induce hyperinflammation and cytokine storms, for example.
A beautiful paper is currently uploaded to the pre-print server OSF PREPRINTS (submitted in November 2021) entitled: “Differences in Vaccine and SARS-CoV-2 Replication Derived mRNA: Implications for Cell Biology and Future Disease”.2 This paper looks at the differences between SARS-nCoV-2 viral spike and the modified mRNA injection spike and asks the question: is the modified spike as, even more pathogenic, than the SARS spike? Basically.
The authors rightly point out in the conclusion that transparency and quality control are necessary aspects of any biological roll out into humans. They penned an extremely well-researched and articulated paper based on the potential devastating effects of codon optimization of the spike protein currently being injected into everyone, but that they are not able to publish it. For some reason. The ‘reasons’ being given to them sound very familiar to the ‘reason’ that Peter and I received in the context of our Myocarditis paper that is still in limbo.
Public and transparent quality control of these often-mandated injections are required. This should include sequence verification and quality control of the various lots and evidence of the proteins these mRNA express in patients.
Another very interesting topic of concern raised in this paper is the following: there is a superantigen residing in the SARS-nCoV-2 spike protein. Was it put there? Before I proceed into this Substack further, I highly recommend reading Kevin, Peter and Anthony’s paper and if you’re interested, Tamara Ugolini interviewed Kevin on this subject matter recently and you can watch that here.
The paper I would like to Substack today is one in their reference list entitled: “Superantigenic character of an insert unique to SARS-CoV-2 spike supported by skewed TCR repertoire in patients with hyperinflammation”.3 The title really says it all. There’s a superantigen in the spike protein and it was inserted.
I find it hard to believe that I only found this paper today. Everyone should be talking about this. What they show is that there is a short sequence of amino acids (epitope) in the SARS-nCoV-2 spike protein that potentially mediates high-affinity, non-specific binding to the T Cell Receptor (TCR) to induce massive non-specific activation of the immune system - cytokine storms and such leading to things like death. This short sequence represents a sequence found in Staphylococcal enterotoxin B (SEB)4 and is called a superantigen due to the above-mentioned properties.
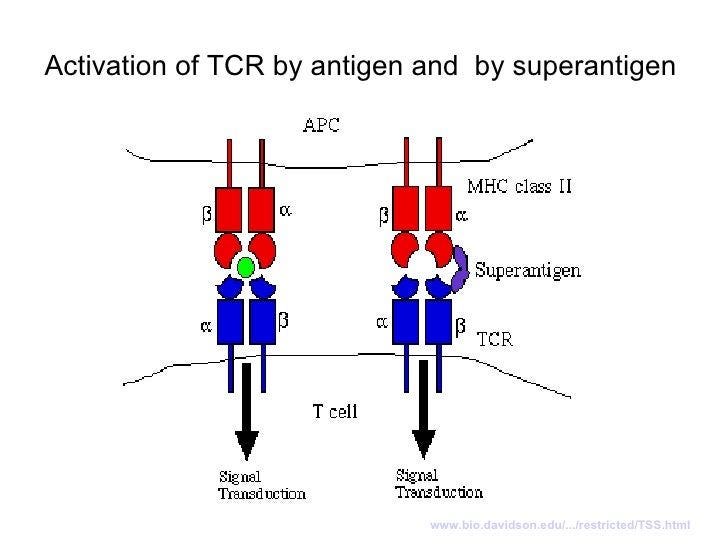
As all of my amazing readers already know, the TCR binds a molecule called an MHC molecule (in this case MHC class II) found on antigen-presenting cells. Good name, eh? In non-superantigen situations, the interaction between the TCR and the MHC-II molecule is much like a sampling situation - if the TCR binds with high affinity to the coupled molecule and specific antigen, signal transduction occurs and the cell goes to work doing what it does! This binding and recognition leads to activation of ~0.01% of T cells. This is in the case of ‘conventional’ antigens. Superantigens, on the other hand, only need to be recognized by the V-beta portion of the TCR (instead of all the other bits) to stimulate 20%! of all T cells!

Basically what these toxins do is remove the necessary specificity component to enable the activation of the T cell, and subsequent effects, to occur via the engaged TCR:MHC-II:antigen interaction, where it normally wouldn’t. All of the functions of that T cell will subsequently ensue and that includes production of massive amounts of cytokines such as IL-2 (mediates tolerance and control of immunity itself and extremely importantly, promotes differentiation of T cells to Treg phenotype) and IFN-gamma (a potent activator of macrophages and inducer of MHC-II molecules). Remember also that antigen-presenting cells such as macrophages produce IL-6 (where have I heard that before?) in response to specific microbial molecules. And the cycle continues.
Onto the paper. Now that you all have amazingly strong backgrounds in immunology and biochemistry, you’ll easily understand why it’s frightening that these authors found that “SARS-CoV-2 Spike Harbors a High-Affinity Site for TCR β-Chain Binding, which Contains An Insertion, PRRA, Unique to SARS2”. That PRRA is the furin-cleavage site that was put there (in my very strong opinion) and is essential to pathogenesis.5 It is not found in SARS-nCoV or in any other coronavirus lineage related.
I need to do something here. A beautifully written hit-piece on the now infamous Pradhan paper has an illustration of an alignment for the 4rth insert discussed in the paper as follows. Let’s see this backfire.

The problem that Houston (and the rest of us) seems to have, has been published in the peer-reviewed literature. One such publication is entitled: “SARS-CoV-2's claimed natural origin is undermined by issues with genome sequences of its relative strains: Coronavirus sequences RaTG13, MP789 and RmYN02 raise multiple questions to be critically addressed by the scientific community”.6
Now, besides hit-piece guy’s typo (GNTS should read QNTS), which I can forgive, he seems to have painted himself into an ironic corner here. The only sequences listed above that include the PRRA sequence (as part of the suggested QTNSPRRA insert), are the variations of the SARS-nCoV-2 (the top 3 sequences). Now, it might just be me, but that seems to prove that this sequence of 4 amino acids that comprises the furin cleavage site (PRRA), which, by the way enhances the infectiousness of the virus by making it more replication competent and virulent (see reference 3), could not have arisen ‘by accident’ or ‘naturally’. We’re talking about 4 (8 actually) sequentially residing amino acids: not 1 or 2. This is not to say that perhaps by some lightning-striking-the-same-place-4-times-in-1-second chance that it didn’t just ‘happen’ to find itself inserted into the SARS-nCoV-2 spike protein in a site that is exceptionally accessible as a potential binding site, but let me just say, alrightttttyyyyyyyyyyy thennnnnnnnnnnnn.
I wanted to make sure everyone knew the relevance and importance of the enhanced destructiveness role of these 4 little amino acids. It makes this spike thing so much worse and you should know that in the referenced paper #3, the version of spike with a PRRA mutant not only attenuated disease in the hamsters they were studying, but yielded robust protection from SARS-CoV-2 re-challenge.
Let’s have a look at the potential superantigen sequence, shall we?
In summary, the authors use computational modeling to show that the SARS-CoV-2 spike contains both sequence and structure motifs that look a whole lot like a superantigen in SEB that may directly bind T cell receptors. They state that the SARS-nCoV-2 spike epitope TNSPRRAR may form a putatively superantigenic core and may not require the participation of the neighboring amino acids. They base this on the binding site properties of the superantigen and the PRRA site: they are very similar sequentially and structurally.
Significantly, the structure of the SARS-CoV-2 S SAg-like segment and that of SEB peptide also exhibit a remarkable similarity: A salt bridge (E159−K152 in SEB and E661−R685 in SARS-CoV-2 S) stabilizes both structural motifs; the relative orientations of three positively charged residues (K152, K153, and K154 in SEB and R682, R683, and R685 in SARS-CoV-2 S) are maintained; and an asparagine (N151 in SEB, N679 in SARS-CoV-2) completes this motif. All three features are absent in SARS1 S. A β-hairpin that apparently serves as a scaffold is conserved in all three spikes, and we observe a pair of cysteines that may potentially form a disulfide bond in SARS-Cov-2 and SARS1 spikes (C662−C671 and C648−C657, respectively).
Now without getting too technical again (too late I guess), it is safe to say that the potential is clear and present for the spike to have a superantigenic site based on these studies. Based on this potential, once again, the precautionary principle dictates that we stop the nonsense of injecting spike protein derived directly from the SARS-nCoV-2 spike into people until we do some science.
Problem is, the science is being done in spite of us. We are the science experiment. Are you ok with that? Psst, the furin cleavage site (and superantigen) is in both the Moderna and Pfizer sequences according to this document entitled “Assemblies of putative SARS-CoV2-spike-encoding mRNA sequences for vaccines BNT-162b2 and mRNA-1273”.

Here are VAERS reports of Toxic Shock Syndrome and Multisystem Inflammatory Syndrome in Children (MIS-C) which are known to be caused by pathogenic superantigens stimulating excessive adaptive immune system activation (T cells) according to the authors and well, everyone. No URF considered. There are 509 reports in VAERS of TSS and MISC in VAERS, so far.

By the way, it has been proposed and published, in fact, that there is a connection between superantigens and sepsis in the context of COVID-19.7 Here are the VAERS sepsis reports. No URF considered. There are 3,999 reports as of July 8, 2022 in VAERS.

Amormino C, Tedeschi V, Paldino G, Arcieri S, Fiorillo MT, Paiardini A, Tuosto L, Kunkl M. SARS-CoV-2 Spike Does Not Possess Intrinsic Superantigen-like Inflammatory Activity. Cells. 2022 Aug 15;11(16):2526. doi: 10.3390/cells11162526. PMID: 36010602; PMCID: PMC9406418.
McKernan, K., Kyriakopoulos, A. M., & McCullough, P. A. (2021, November 25). Differences in Vaccine and SARS-CoV-2 Replication Derived mRNA: Implications for Cell Biology and Future Disease. https://doi.org/10.31219/osf.io/bcsa6.
Cheng MH, Zhang S, Porritt RA, Noval Rivas M, Paschold L, Willscher E, Binder M, Arditi M, Bahar I. Superantigenic character of an insert unique to SARS-CoV-2 spike supported by skewed TCR repertoire in patients with hyperinflammation. Proc Natl Acad Sci U S A. 2020 Oct 13;117(41):25254-25262. doi: 10.1073/pnas.2010722117. Epub 2020 Sep 28. PMID: 32989130; PMCID: PMC7568239.
https://www.health.pa.gov/topics/Documents/Diseases%20and%20Conditions/Staphylococcal%20Enterotoxin%20B%20.pdf
Johnson BA, Xie X, Kalveram B, Lokugamage KG, Muruato A, Zou J, Zhang X, Juelich T, Smith JK, Zhang L, Bopp N, Schindewolf C, Vu M, Vanderheiden A, Swetnam D, Plante JA, Aguilar P, Plante KS, Lee B, Weaver SC, Suthar MS, Routh AL, Ren P, Ku Z, An Z, Debbink K, Shi PY, Freiberg AN, Menachery VD. Furin Cleavage Site Is Key to SARS-CoV-2 Pathogenesis. bioRxiv [Preprint]. 2020 Aug 26:2020.08.26.268854. doi: 10.1101/2020.08.26.268854. PMID: 32869021; PMCID: PMC7457603.
Deigin Y, Segreto R. SARS-CoV-2's claimed natural origin is undermined by issues with genome sequences of its relative strains: Coronavirus sequences RaTG13, MP789 and RmYN02 raise multiple questions to be critically addressed by the scientific community. Bioessays. 2021;43(7):e2100015. doi:10.1002/bies.202100015.
Scaglioni V, Soriano ER. Are superantigens the cause of cytokine storm and viral sepsis in severe COVID-19? Observations and hypothesis. Scand J Immunol. 2020;92(6):e12944. doi:10.1111/sji.12944.


As Michael Yeadon writes, no way are these "vaccines" accidental in their pathology. It has been long known that the spike IS the toxin that is causing all the damage of the covid virus. Yet all 3 companies decided to use spike as the TARGET.
And they recommended it for EVERYONE, including pregnant women. That has NEVER happened before.
Read watch Yeadon. He explains it concisely.
You are a ROCK STAR Jessica. Looking back, we will not be ignorant of the evidence of criminality, mens rea, and the amount of deviousness that went into engineering this bio weapon.
So, to put this in laymen's terms, we are screwed. We need to stop these injections but the money and power has so overcome all ethics and morals that it seems unstoppable. What will it take to stop it? I assume it has to be a complication in one of our elite rulers or in their families. Celebrity complications were not enough. We just make up a new category of sudden death of unknown causes.