Evidence of connection between Severe Adverse Events and mRNA degradation
Is it possible that as the mRNA degrades the risk of severe adverse events decreases?
There’s a new paper in town that is more akin to a thesis. It is entitled: “Immune imprinting, breadth of variant recognition and germinal center response in human SARS-CoV-2 infection and vaccination Cell”.1 It’s a high-level piece of work and has caught the eye of every serious scientist that I know including Robert Malone, Stephanie Seneff, Kevin Mckernan, Peter McCullough and myself. We discussed it at length last night on our weekly call to discuss COVID-19 insanities. Robert has penned an excellent review on his Substack (like he needs a call-out from me, right?)
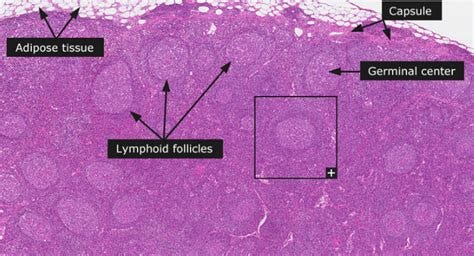
The gist of it is this: Spike antigen and mRNA from the injections hang around for at least 60 days in germinal centers of lymph nodes.
Germinal centers2 of the lymph nodes are dynamic specialized places where high affinity antibodies get manufactured via B cell clonal expansion (more shoes please), somatic hypermutation (making shoes of different sizes) and affinity-based selection (trying on different shoes to see which ones fit you).
This is what you want to have happen in a natural response to a foreign antigen. But this becomes problematic for obvious reasons when foreign antigen or protein is being self-produced in vast quantities.
But this is not the focus of this article. I just wanted to say that there is evidence now in the literature that the mRNA of this spike protein sticks around in germinal centers. It’s stable and it’s functional.
I have an ongoing hypothesis that the severe adverse event reports in VAERS (and Yellowcard and Eudra) might be related to the state of the mRNA templates delivered to the person via injection by (Lipid Nano Particles) LNPs and not just necessarily the dose. When a person gets injected with the Pfizer product, they are meant to get injected with 30 ug of mRNA as part of the LNP ‘mix’. But my question is, even if the dose is correct, what is the state of the mRNA? If the mRNA templates are not complete, they will not be translatable to complete spike proteins. This could either be a good thing or a bad thing in terms of the effects. We have no idea what the effects of generation of massive quantities of complete spike proteins would be on human physiology (although we are beginning to find out), let alone the effects of incomplete spike proteins. This is documented by Pfizer as an unknown.
The potential safety risks associated with truncated RNA isoforms (‘functionally similar’ versions) are not addressed.
This scared the shit out of me when I read it. The true percentages of the mRNA integrity on a subset of Pfizer vax lots reveal that they barely have 55% integrity.
Since we don’t know the effects of ‘low’ mRNA integrity on translation products, we can’t know yet if the effects would be more or less detrimental. The full spike protein is proving to be quite detrimental to human physiology so is it possible that incomplete versions may result is lessened pathologies? Is it also possible that this could explain the differences between people with regard to adverse event experiences? I mean, not everyone is succumbing, and my thoughts go to mRNA integrity and lack of mRNA altogether. The question remains on the subject of ‘placebo’ with these mRNA COVID-19 products - are they saline or are they empty LNPs?
I have what might be a confirmation that the truncated versions of the mRNAs may lead to fewer mal effects, at least in the short term. This would make me feel better from one point of view, but not so good from the point of view of what the mRNA → spike protein is actually doing in the full-template form. If one plots the number of severe adverse events (SAEs) reported to the Domestic data set in VAERS in the context of the COVID-19 Pfizer products against the time between the product manufacturing date of each VAX LOT and the injection date, you see the following plot. By the way, this pattern is lost when one includes the foreign data.

This plot includes Pfizer severe adverse event reports from the Domestic VAERS data set with cleaned VAX LOT data. Let’s ask a question.
Q: Is there a relationship between the time to injection from manufacturing date and severe injury reports?
The above plot appears to demonstrate that the reports of SAEs are highest following injection date ~1 month after manufacturing date. Since I don’t have a lot of data pertaining to injections given with fresh products (<1 month), I suspect that if I did, that the trajectory would look more like the orange line in the following plot. Sorry for the noise in the blue trajectory.
Many assumptions would have to be made here for this to qualify as theoretical evidence. For example, one would have to assume that the mRNA/LNP integrity would degrade according to time. This isn’t a hard assumption to make and accept since this is pretty much what an expiration date is. One would also have to assume that degraded mRNA would manifest as non-’effective’ with regard to causing damage.
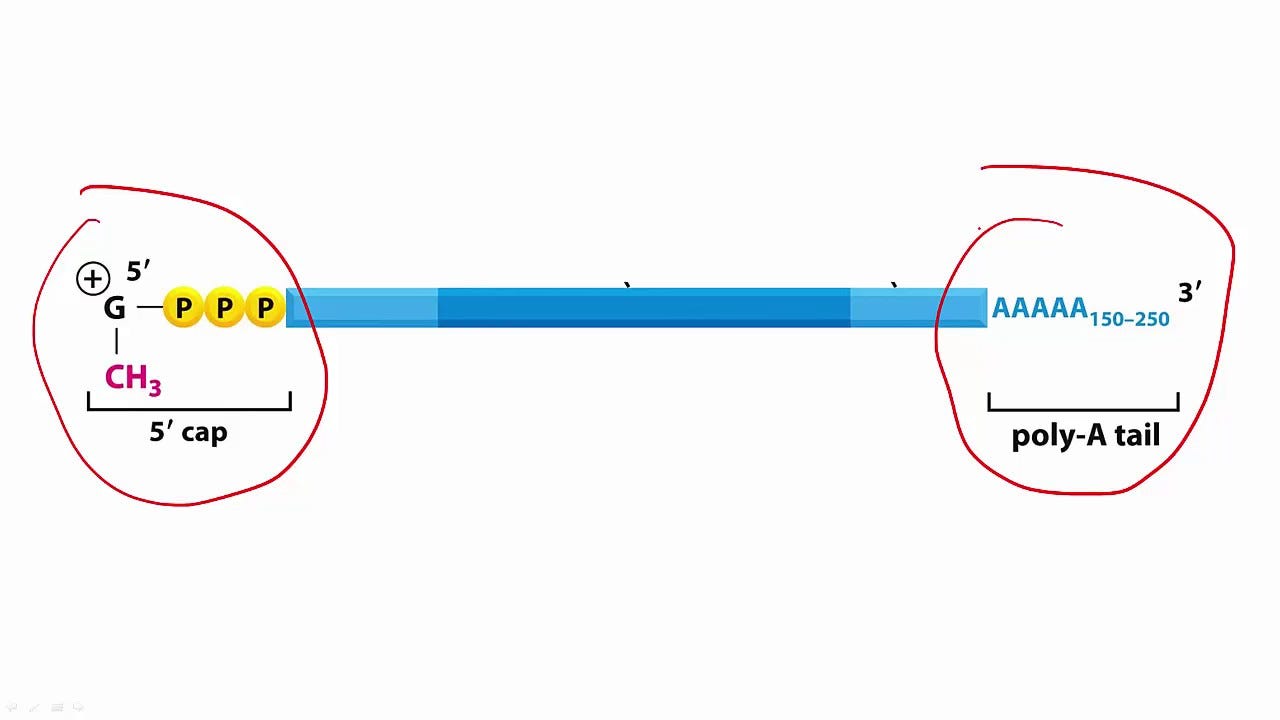
Insertion: One must wonder what degraded mRNA actually means at this point. ie: Where is the mRNA truncated and what effects do different truncations have? For example, if the truncation occurs at the poly-A tail end, it is still possible for the mRNA to be translated to some other protein? What would the effect of these novel proteins be? Are these proteins mis-folded? Are they akin to prions?
For those who don’t know, mRNAs are book-ended between a ‘cap’ and a ‘tail’. The cap, or 5’ cap (ribosome moves along the mRNA in the 5' (‘left’)-to-3' (‘right’) direction) and the 3’ poly-A tail protect the mRNA transcript (the middle bit in darker blue below) and aid in nuclear export and translation. What are the effects of degradation of the poly-A tail with regard to translation once injected? Are there effects at all or is the mRNA not translatable? What of the untranslated regions (UTRs)? What about the effect of translation fidelity due to the pseudouridine replacements? What are the effects of the humanized codon optimization? To what extent does the codon optimization - with GC content differences - cause protein folding impairment upon translation (look up ribosomal pausing3)? What of the mutated forms of S1 in the injected? There are far too many unanswered questions.
I am going to pause for now. I need to do some more thinking and reading and some more VAERS seeking.
I would also love comments on this.
Published: January 24, 2022. DOI: https://doi.org/10.1016/j.cell.2022.01.018
Victora GD, Nussenzweig MC. Germinal centers. Annu Rev Immunol. 2012;30:429-57. doi: 10.1146/annurev-immunol-020711-075032. Epub 2012 Jan 3. PMID: 22224772.
Martine A Collart, Benjamin Weiss, Ribosome pausing, a dangerous necessity for co-translational events, Nucleic Acids Research, Volume 48, Issue 3, 20 February 2020, Pages 1043–1055, https://doi.org/10.1093/nar/gkz763





Thank you Ms. Rose. I must confess that this is over my head, so any comment from me would be the peanut gallery.
I can safely say however, that we are blessed to have you on the case and you and your peers will get to the bottom of it.
I will be following along and continue to learn from your work.
If most injections are also ~30 days after manufacture, product expiration would be redundant in terms of the resulting AE plot and real world impact. So we would need the injections by date after manufacture plot to see if there are any people actually being protected by hypothetical product expiry.
On the other hand if manufacturer, cold chain, and dosage inconsistency determine functioning mRNA payload, it’s pure Russian Roulette. I think Bhargava recently mentioned LNP bubble size as a potential payload randomizer. But VAERS is full of dilution/injection errors, including for the 5-11 product.
But the Röltgen, K. et al. paper suggests consistent payload at least in the sense that no one is getting “no spike.” 96% of recipients have detectable spike in plasma on day 1-2 and 63% at day 7. Similar results for mRNA and spike in the germ centers, with lots of hits still on day 37 post 2nd-dose and few after. So it seems like everyone who gets the shot is getting a lot of intact mRNA. The variability is in how long it sticks around.
Not discussed much, the paper also finds aberrantly cross-protective antibody enrichment against the RBD vs non-mRNA shots or infection with “Wuhan” strain (fig 4), as in binding against variants / binding against Wuhan ratio was much more even for Pfizer recipients. Even though the mRNA supposedly codes for the Wuhan spike. This suggests there is variability in the output, perhaps lots of mutant spike production, though it could also be some weird influence on APC epitope selection, maybe from the pseudouridine, maybe from the “slow release” factor of mRNA continuing to become available to APCs for weeks after dosage. Malone says the results suggest the mRNA is lingering in the immune cells but I think it reflects delayed capture from transfected non-immune cells.
Perhaps someone should have tested how these things work in a lab or something.